M. Klika Škopić, K. Götte, C. Gramse, M. Dieter, S. Pospich, S. Raunser, R. Weberskirch, A. Brunschweiger
https://doi.org/10.1021/jacs.9b05696
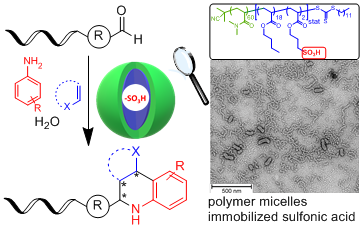
DNA-encoded libraries are increasingly used of drug identification. However, the reactivity of the DNA imposes limitations on the choice of chemical methods for encoded library synthesis. Amphiphilic block copolymers covalently functionalized with sulfonic acid moieties in the lipophilic portion assemble in water and locate the Brønsted catalyst in micelles. These acid nanoreactors enabled the reaction of DNA-conjugated aldehydes to diverse substituted tetrahydroquinolines and aminoimidazopyridines by Povarov and Groebke–Blackburn–Bienaymé reactions, respectively.