Paper Highlights
The reverse transcription signature of N-1-methyladenosine in RNA-Seq is sequence dependent
Ralf Hauenschild, Lyudmil Tserovski, Katharina Schmid, Kathrin Thüring, Marie-Luise Winz, Sunny Sharma, Karl-Dieter Entian, Ludivine Wacheul, Denis L. J. Lafontaine, James Anderson, Juan Alfonzo, Andreas Hildebrandt, Andres Jäschke, Yuri Motorin, Mark Helm
Nucleic Acids Res. 2015, DOI:10.1093/nar/gkv895
Bei der reversen Transkription hinterlässt 1-Methyladenosin Spuren in der cDNA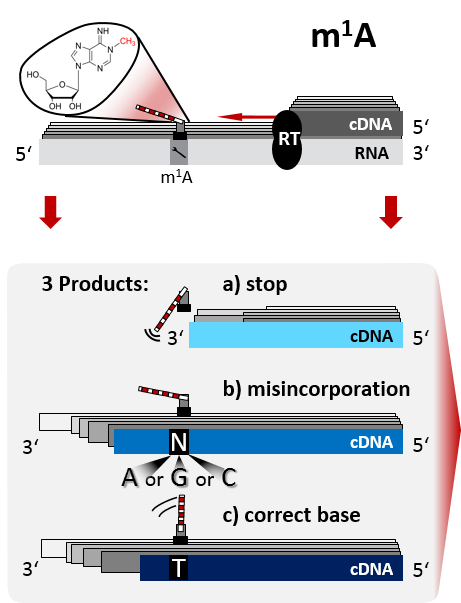
Extraordinary thermal stability of an oligodeoxynucleotide octamer constructed from alternating 7-deaza-7-iodo guanine and 5-iodocytosine base pairs – DNA duplex stabilization by halogen bonds?
Natalya Ramzaeva, Henning Eickmeier, Helmut Rosemeyer
Chem. Biodiversity 2014, DOI:10.1002/cbdv.201300300
A reinvestigation of the published X-ray crystal-structure analyses of 7-halogenated (Br, I) 8-aza-7-deaza-2’-deoxyguanosines Br7c7z8Gd; 1a and I7c7z8Gd, 1b, as well as of the structurally related 7-deaza-7-iodo-2’-deoxy-ß-D-ribofuranosyladenine (ß-I7c7Ad; 2-6e in Table 1) and its α-D-anomer (α -I7c7Ad; 3) clearly revealed the existence of halogen bonds between corresponding halogen substituents and the adjacent N(3)-atoms of neighboring nucleoside molecules within the single crystals. These halogen bonds can be rationalized by the presence of a region of positive electrostatic potential, the σ-hole, on the outermost portion the halogen’s surface, while the three unshared pairs of electrons produce a belt of negative electrostatic potential around the central part of the halogen substituent. The N(3) atoms of the halogenated nucleosides carry a partial negative charge. This novel type of bonding between nucleosides was tentatively used to explain the extraordinary high stability of oligodeoxynucleotides constructed from halogenated nucleotide building blocks.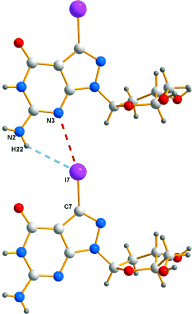
‘Yeast Mail’: A novel saccharomyces application (NSA) to encrypt messages
Helmut Rosemeyer, Achim Paululat, Jürgen J. Heinisch
Chem. Biodiversity 2014, DOI:10.1002/cbdv.201400160
The universal genetic code is used by all life forms to encode biological information. It can also be used to encrypt semantic messages and convey them within organisms without anyone but the sender and recipient knowing, i.e, as a means of steganography. Several theoretical, but comparatively few experimental, approaches have been dedicated to this subject, so far. Here, we describe an experimental system to stably integrate encrypted messages within the yeast genome using a polymerase chain reaction (PCR)-based, one-step homologous recombination system. Thus, DNA sequences encoding alphabetical and/or numerical information will be inherited by yeast propagation and can be sent in the form of dried yeast. Moreover, due to the availability of triple shuttle vectors, Saccharomyces cerevisiae can also be used as an intermediate construction device for transfer of information to either Drosophila or mammalian cells as steganographic containers. Besides its classical use in alcoholic fermentation and its modern use for heterologous gene expression, we here show that baker’s yeast can thus be employed in a novel Saccharomyces application (NSA) as a simple steganographic container to hide and convey messages.
RNA interference controlled by light of variable wavelength
Andreas Meyer, Andriy Mokhir
Angew. Chem. Int. Ed. 2014, DOI:10.1002/anie.201405885
Known “caged“ siRNAs are activated by UV light, which can be toxic to cells. An siRNAs that can be activated by red light under mild conditions is reported. The uncaging is mediated by 1O2 photo-generated by a photosensitizer, which is attached to the 3’-terminus of the lagging strand. The 5’-terminus of the guide strand is alkylated with a 9-anthracenyl residue.
Endless: A purine-binding RNA motif that can be expressed in cells
Christoph Kröner, Martin Thunemann, Sven Vollmer, Manuela Kinzer, Robert Feil, Clemens Richert
Angew. Chem. Int. Ed. 2014, DOI:10.1002/anie.201403579
A de novo designed RNA motif binds cGMP with low micromolar affinity and, when inducibly expressed in cells, acts as sink for this second messenger.
Influence of the absolute configuration of NPE-caged cytosine on DNA single base pair stability
H. S. Steinert, F. Schäfer, H. R. A. Jonker, A. Heckel, H. Schwalbe
Angew. Chem. Int. Ed. 2014, DOI:10.1002/anie.201307852
Photolabile protecting groups are a versatile tool to trigger reactions by light irradiation. In this study, we have investigated the influence of the absolute configuration of the 1-(2-nitrophenyl)ethyl (NPE) cage group on a 15-base-pair duplex DNA. Using UV melting, we determined the global stability of the unmodified and the selectively (S)- and (R)-NPE-modified DNA sequences, respectively. We observe a differently destabilizing effect for the two NPE stereoisomers on the global stability. Analysis of the temperature dependence of imino proton exchange rates measured by NMR spectroscopy reveals that this effect can be attributed to decreased base pair stabilities of the caged and the 3′-neighbouring base pair, respectively. Furthermore, our NMR based structural models of the modified duplexes provide a structural basis for the distinct effect of the (S)- and the (R)-NPE group.