Paper Highlights
On-oligonucleotide olefin metathesis in water
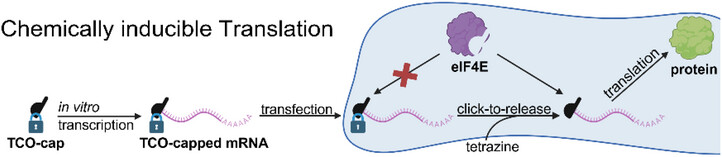
Small-Molecule Activation of mRNA Translation by Click-to-Release Reaction in Cells
T. Vosman, F. Burba, M. Sumser, M. Vrabel, H. Mikula, A. Rentmeister
Angew. Chem. Int. Ed. 2026, 65, e24223
DOI: 10.1002/anie.202524223
A new method to control mRNA activity using bioorthogonal click-to-release reactions is presented. The 5′ cap is modified with a trans-cyclooctene (TCO) that quickly reacts with hydroxyaryl-tetrazines and efficiently releases the native cap 0. TCO-capped mRNAs are initially translationally inactive but can be activated with non-toxic, cell-permeable tetrazines. This strategy offers a versatile way to regulate mRNA translation in eukaryotic cells.
ProTides for Antiviral Activity Beyond Liver Cells
F. Goebel, A. Humboldt, H. Kim, C. J. M. Schölzel, R. Bartenschlager, C. Richert
Chem. Eur. J. 2026
DOI: 10.1002/chem.70901
A strategy for obtaining prodrugs of antiviral nucleotides with broad tissue activity is presented that relies on cycloalkyl or cycloalkylalkyl esters, improving uptake and esterase cleavage, and producing nanomolar inhibitors in kidney, colon, and lung cells.

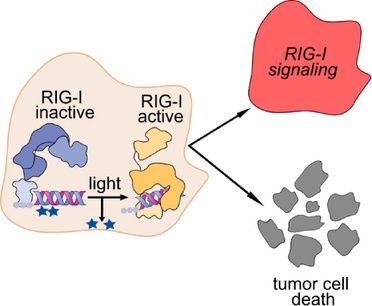
Immunoengineering of a Photocaged 5´-triphosphate Oligoribonucleotide Ligand for Spatiotemporal Control of RIG-I Activation in Cancer
S. A. Lewash, V. R. McKenney, C. Wuebben, J. Ludwig, R. Hosni, D. Radzey, M. I. Toma, E. Bartok, M. Schlee, T. Zillinger, A. Heckel, G. Hartmann
Angew. Chem. Int. Ed. 2025 , 64 , e202423321
DOI: 10.1002/anie.202423321
Photochemical control of oligonucleotides bears great potential for the spatio-temporal control of therapeutic targets, such as immune sensing receptors. Retinoic acid-inducible gene I (RIG-I) is a cytoplasmic receptor of the innate immune system that triggers antiviral responses upon detection of viral RNA. RIG-I can be specifically activated by short double-stranded (ds) RNA with a blunt 5′ end bearing a triphosphate, mimicking nascent viral transcripts. Tumor cells are specifically sensitive to RIG-I-induced cell death. Here we developed a potent oligonucleotide ligand for spatiotemporally controlled activation of RIG-I by light exposure. Through structural considerations and functional studies we identified a combination of two nucleoside positions in a RIG-I oligonucleotide ligand for which the substitution of both respective 2′-hydroxy groups of the ribose by photolabile protecting groups (2′-photocages) resulted in a complete loss of RIG-I ligand activity, whereas photocaging the individual positions was not sufficient to turn off RIG-I. Light exposure fully restored RIG-I activation by the photocaged RIG-I ligand, enabling light-controlled RIG-I-mediated cell death of human cancer cells which had internalized the photocaged RIG-I ligand prior to light exposure. This novel photoactivatable RIG-I oligonucleotide ligand may be applicable for precise light-controlled induction of tumor cell death in superficial cancer such as melanoma.
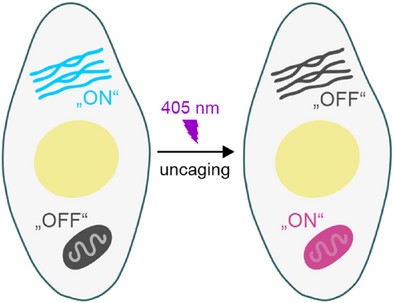
Wash-Free Multi-Target Super-Resolution Microscopy With Photocaged DNA Labels
Nina Kaltenschnee, Marina S. Dietz, Laurell F. Kessler, Yunqing Li, Janik Kaufmann, Alexander Heckel, Mike Heilemann
Angew. Chem. Int. Ed. 2026, 65, e26137
DOI: 10.1002/anie.202526137
Super-resolution microscopy with DNA-fluorophore labels is primed for multi-target imaging of cell biological samples. However, direct interaction with the sample is required to exchange or add DNA-fluorophore labels in each imaging round, which can impair the accuracy of the imaging data at the nanometer scale. To bypass this requirement, we introduce PhotoPAINT, a wash-free method that employs DNA oligonucleotides equipped with photocaging groups. Irradiation with light removes these photo-modulatable groups and changes the hybridization properties of DNA labels, enabling light-modulated targeting. We demonstrate this concept by imaging various cellular targets with confocal microscopy, single-molecule localization microscopy (SMLM), and stimulated emission depletion (STED) microscopy.
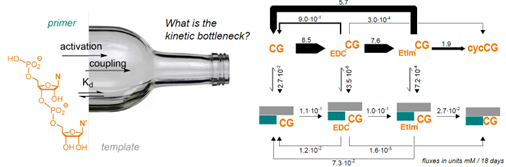
A quantitative model of enzyme-free copying of RNA with dimers
L. Burger, F. Welsch, E. Kervio, M. Henker, G. Leveau, U. Gerland, C. Richert
Nucleic Acids Res. 2025, 53, gkaf987.
https://doi.org/10.1093/nar/gkaf987
Seeing all relevant molecular processes involved in extending a template-bound RNA strand by 12 bases in one run, starting from unactivated dinucleotides in dilute aqueous buffer in the cold.